| Highly efficient and specific | Any gene |
| Value for money | Time-saving, detailed design |
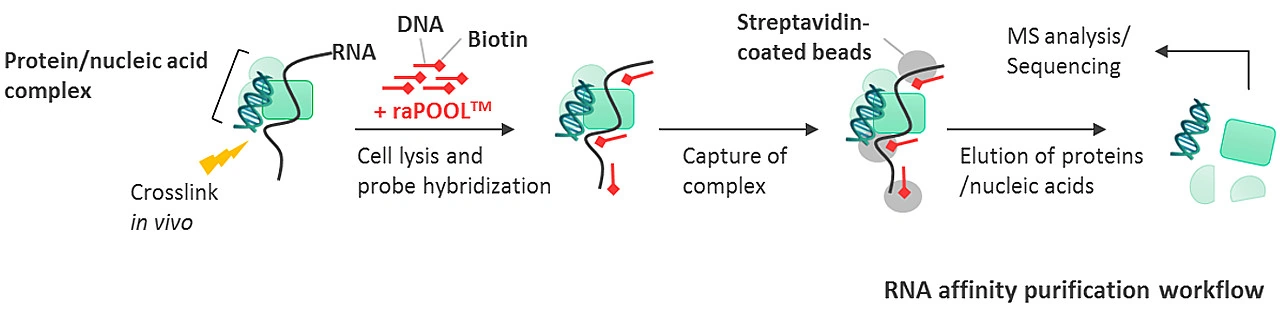
Nucleic acid and protein interactions are preserved with a cross-linking reagent, followed by lysis and sonication to shear nucleic acids to sizes amenable for pulldown. The raPOOL is added to lysates and hybridizes with the RNA of interest. Addition of streptavidin-coated magnetic beads allow isolation of the raPOOL-bound complex through the high affinity biotin-streptavidin interaction. The complexes are then disrupted and individual components analysed by various methods: western blotting/mass spectrometry (for proteins), sequencing/northern and southern blots/PCR detection (for nucleic acids).
Please use the form below to request for a quote or ask for more information about raPOOL.